This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
proofread
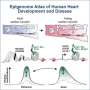
New tool allows for gene suppression in mouse heart muscle cells using CRISPRi

An innovative tool for the targeted modification of gene activity in heart muscle cells could establish itself as a standard method for research into cardiovascular diseases.
Dr. Patrick Laurette and his colleagues at the German Center for Cardiovascular Research (DZHK), led by Prof. Ralf Gilsbach, have successfully reduced the activity of individual genes in mouse heart muscle cells using the CRISPRi system.
This technology allows for the temporary suppression of gene expression without altering the genetic sequence. It thus avoids the potential risks associated with direct intervention in the genome. The study is published in Circulation Research.
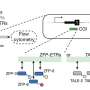
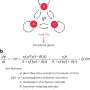
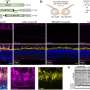
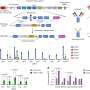
CRISPRi is based on the CRISPR-Cas genome editing system but lacks the ability to cut DNA, a so-called "dead CAS" (dCAS) and is fused to a KRAB repressor domain. Using a short RNA as a molecular guide dCas9 can bind to specific DNA sequences. As a result, the target region in the genome is epigenetically silenced, and a gene can no longer be read. This blockade is called "epigenetic silencing."
To introduce the CRISPRi system into the heart muscle cells of mice, the researchers used adeno-associated viruses (AAV), which do not integrate into the genome. One challenge was to package the entire system into the limited genomic capacity of the AAVs. The researchers succeeded in doing this by using a particularly small dCAS. Although other viral vectors have more capacity in this respect, they cannot efficiently reach heart muscle cells or integrate into the genome.
Effective blockade in the heart muscle
Dr. Patrick Laurette and his colleagues at Heidelberg University Hospital demonstrated how well epigenetic silencing with the AAV-CRISPRi system works in heart cells for several genes and enhancers. The activity of some genes decreased by up to 95 percent. Enhancers are regulatory elements that can fine-tune gene expression from distant genomic regions.
"The complexity of the mammalian organism is the result of around 1 million regulatory elements. There are between 50,000 and 100,000 of these enhancers in heart muscle cells alone," says Prof. Ralf Gilsbach. They are now focusing on modulating these regulatory elements using their new method to treat diseases such as heart failure or cardiac arrhythmia.
Translational perspective
Gilsbach emphasizes the translational significance of this approach, which makes it possible to specifically influence gene expression in vivo without changing the DNA sequence. The method is also titratable, meaning it can be regulated and its effect can be reversed. Compared to other methods, such as genetic knock-out, in which genes are destroyed, the AAV-CRISPRi system offers a more precise imitation of natural regulatory mechanisms.
Methodically optimized and further developed, the procedure could also be used for human therapy in the long term. "I am convinced that this approach has translational significance, even if it is difficult to predict how quickly development will continue here," says Gilsbach.
There are already numerous companies that are considering AAVs to deliver CRISPR components for therapies. Among other things, they are working on avoiding unwanted antibody reactions. This is because humans have antibodies against AAV and often also against the CRISPR protein derived from bacteria.
More information: P. Laurette et al, In Vivo Silencing of Regulatory Elements Using a Single AAV-CRISPRi Vector, Circulation Research (2023). DOI: 10.1161/CIRCRESAHA.123.323854